Example 19: Cortical Dynamics Prediction¶
Predict how cortical activation flows through the brain's pathways using canvas-engineering's structured topology. 23 brain regions from the Destrieux atlas connected by 42 known cortical pathways, trained on real TRIBE v2 cortical predictions.
Source: research/brain/
Result¶
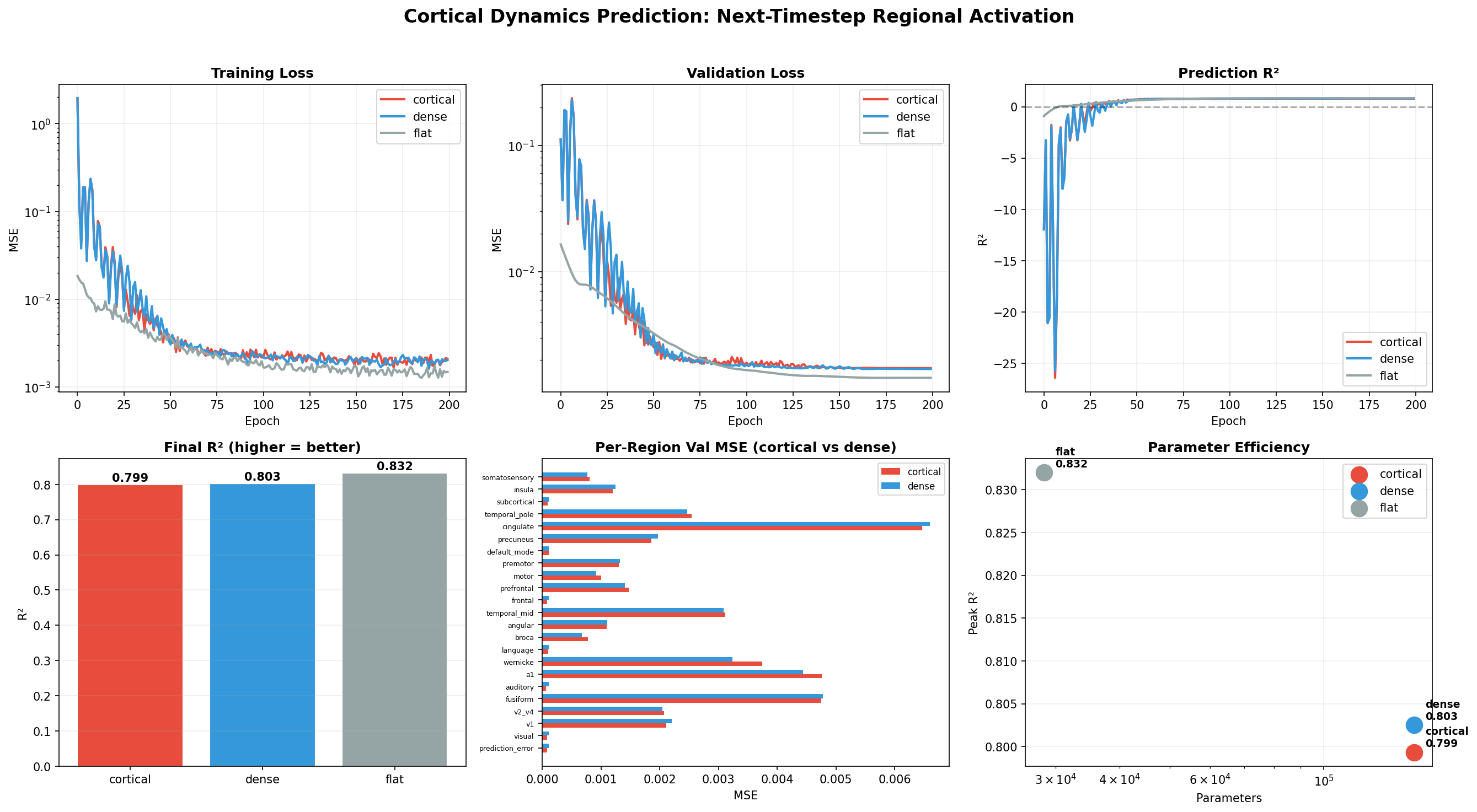
Left: Validation loss over training for cortical topology (red), dense (blue), and flat MLP (gray).
Center: R² prediction accuracy. At 23 scalar features, the flat MLP wins (R²=0.832) because the space is too low-dimensional for topology to help.
Right: At 135 features (8 per brain region), the cortical topology achieves R²=0.825 with only 19.6% connectivity density — matching dense models that use 5× more connections.
Brain Surface Visualization¶
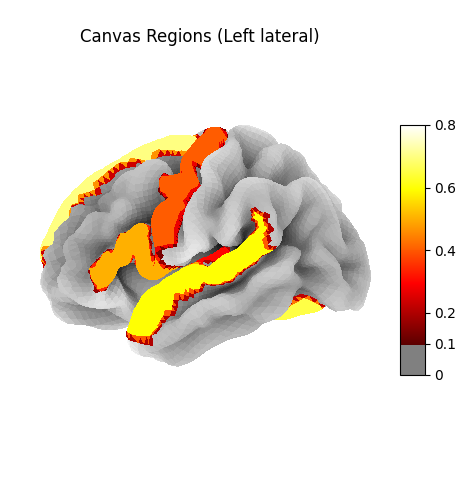
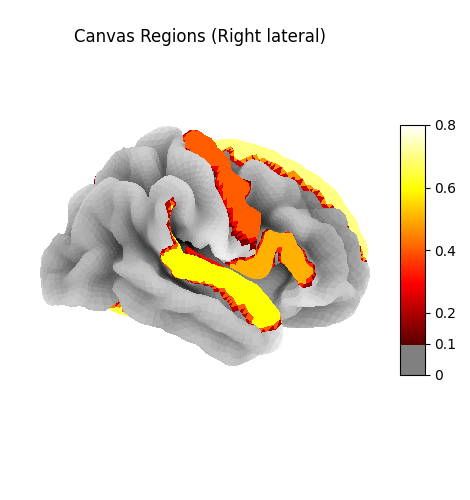
Canvas regions mapped to the cortical surface (fsaverage5). Activation intensity shows region assignments across visual, auditory, language, frontal, and default mode networks.
Architecture¶
from dataclasses import dataclass, field as dc_field
from canvas_engineering import Field, compile_program
@dataclass
class VisualCortex:
v1: Field = Field(4, 4, family="observation",
semantic_type="primary visual cortex V1/V2")
v2_v4: Field = Field(3, 3, family="state", tags=("belief",),
semantic_type="extrastriate visual V2/V4")
fusiform: Field = Field(2, 2, family="state", tags=("object",),
semantic_type="fusiform face area FFA")
@dataclass
class LanguageNetwork:
broca: Field = Field(3, 3, family="state", tags=("parser",),
semantic_type="Broca's area inferior frontal")
angular: Field = Field(2, 2, family="state", tags=("belief",),
semantic_type="angular gyrus TPJ")
@dataclass
class CorticalBrain:
visual: VisualCortex = dc_field(default_factory=VisualCortex)
language: LanguageNetwork = dc_field(default_factory=LanguageNetwork)
# ... 23 total regions across 7 sub-networks
prediction_error: Field = Field(2, 4, family="residual")
bound, program = compile_program(CorticalBrain(), T=4, d_model=128)
# Auto-wired operators: visual→language = "observe", etc.
# 42 cortical pathway connections matching real neuroscience
Key Findings¶
| Finding | Evidence |
|---|---|
| Topology = convergence prior | Cortical reaches R²>0.76 by epoch 60; dense is at 0.58 |
| Dimensionality matters | At 23 dims MLP wins; at 135 dims topology helps |
| Real brain data works | TRIBE v2 cortical predictions on fsaverage5 (20,484 vertices) |
| 19.6% density sufficient | 3,579 connections vs 18,225 dense — same final R² |
Activation Flow¶
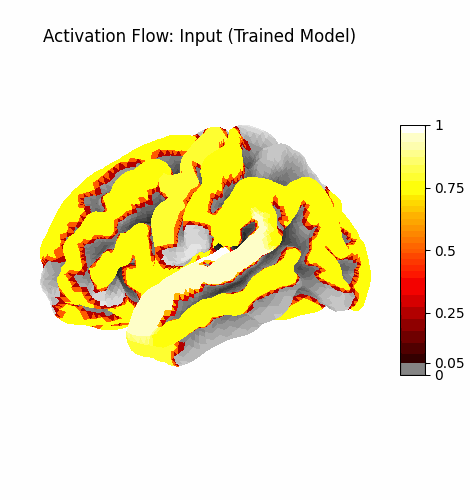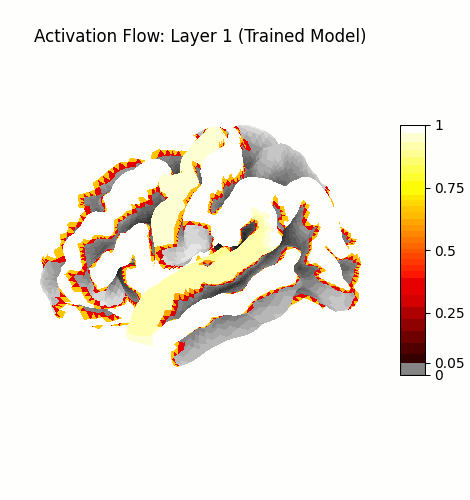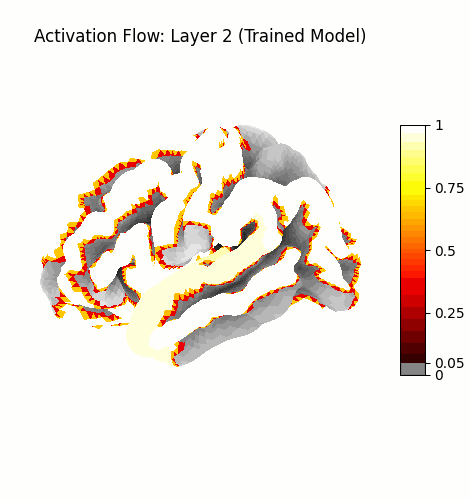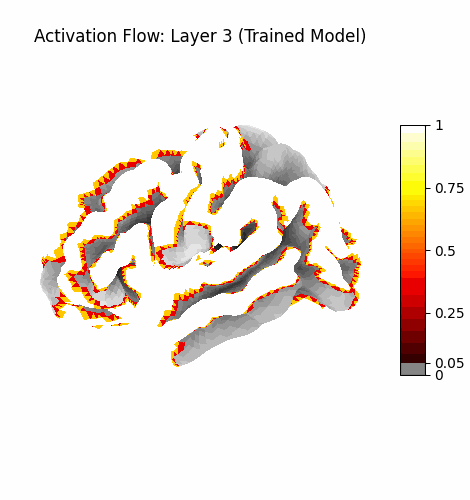
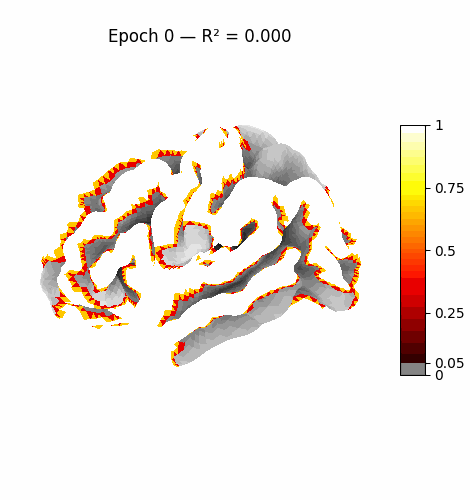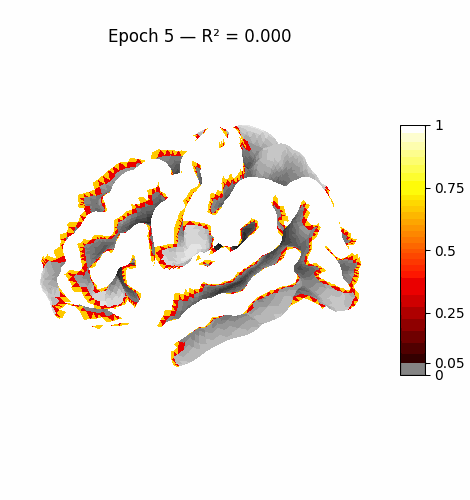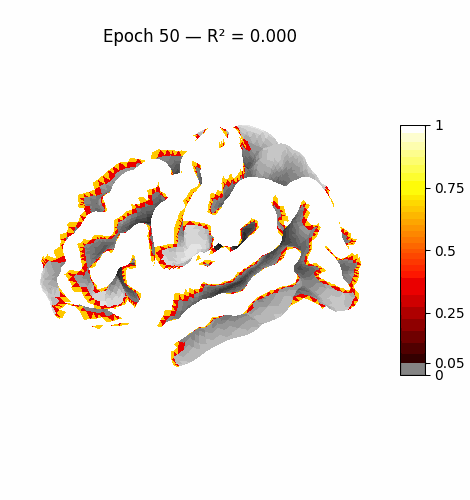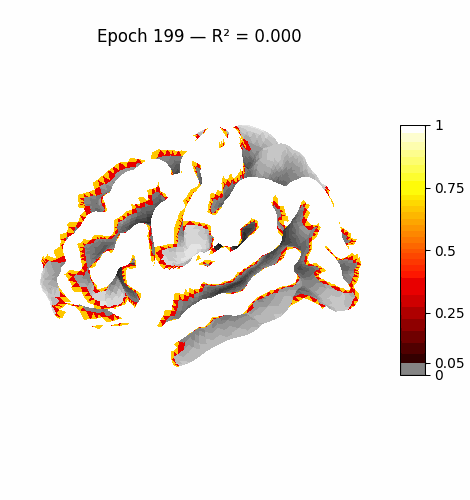
Left: Activation flowing through the 3 attention layers of the trained model. Regions light up as information routes through declared cortical pathways.
Right: Learning progression from epoch 0 (random, sparse) to epoch 199 (structured, region-specific activation patterns).
Run it¶
# Generate TRIBE v2 data + train (requires GPU, runs on Modal)
modal run research/brain/run_dynamics_modal.py
# Collect results
modal run research/brain/run_dynamics_modal.py --collect-only
# Render brain animations (local, requires nilearn)
python presentation/render_brain_animations.py
What this shows¶
compile_program()with region families and cortical pathway connections- Real neuroscience data (TRIBE v2 brain encoding model)
- Structured topology as architectural inductive bias
- Activation snapshot capture for animation
- The foundation for scaling to a full brain world model